top of page

Research
Biomolecules inherently exert their unique physiological functionsthrough a variety of dynamics, such as structural changes trig-gered by variations in the external environment, binding ofsubstrates, and association and dissociation with other pro-teins. Hence, detailed information on various dynamic prop-erties of proteins on the sub-second timescale obtained bydirectly filming them as movies can provide deep insightsand understanding of essential mechanisms of their functionalexpression. The development of various types of bioimagingtechniques based on optical microscopy has enabled us totrack movements of proteins at the single molecular level n vitro and even in vivo and has contributed greatly to theprogress of life science. On the other hand, what is generally being imaged by opticalmicroscopy techniques is not the protein itself, but the changein the locus of the fluorescent regions and the fluorescenceintensity emitted by the labeled molecules bound to the pro-tein. The behavior of the protein is indirectly determined fromthese observations. To understand intuitively proteins’func-tions, a microscopy technique capable to resolve their higherorder structure and high-speed imaging in solution is needed. Our laboratory is developing single molecule dynamics measurement techniques based on high-speed atomic force microscopy (AFM), and is aiming to understand the physical mechanisms of protein function expression by single molecule dynamics imaging.
The following is examples of protein functional dynamics recorded by high-speed atomic force microscopy.

Cellulolytic enzyme

F 1 -ATPase
AFM (Atomic Force Microscopy)
Atomic Force Microscopy (AFM) was invented by G. Binning (IBM Zurich) and C. Quate (Stanford University) in 1986 [1]. G. Binning invented the Scanning Tunneling Microscope[2] in 1982 and was awarded the Nobel Prize in Physics in 1986, but the STM can only observe conductive samples to measure the tunneling current between the conductive needle and sample. On the other hand, since AFM is capable of observing the surface structure of insulator samples with high resolution, AFM has become one of the indispensable instruments for nanoscience and nanotechnology. In AFM, a sharp probe attached to the end of a soft spring (cantilever) is brought into contact with the sample surface, and the mechanical interaction between the probe and the sample is detected by the displacement of the cantilever. The surface structure of sample can be imaged by 2D mapping of the forces acting on the probe. By detecting Coulomb or magnetic forces acting on the metalic or magnetic probe, local electrical.amagnetic information can also be mapped. Also analysis of cantilever oscillation characteristics can measure the local mechanical properties of the sample, providing a high-resolution various physical properties in addition to the surface structure (there are many microscopy techniques derived from STM and AFM, collectively called scanning probe microscopy.). One of advantages of AFM is that it can be used in any operating environment, allowing observation of samples in solution as well as in vacuum and air. Therefore, the development of in-solution AFM and its application to biological samples have been vigorously pursued immediately after its invention, and a wide variety of samples ranging from proteins, nucleic acids, and chromosomes to cells have been imaged [2]. However, since AFM generally requires several tens of seconds to several minutes to acquire a single frame, it has been difficult to observe phenomena involving motion, such as intermolecular interactions, assembly and structural changes of molecules.
1. G. Binnig, CF Quate, & Ch. Gerber, "Atomic Force Microscope", Phys. Rev. Lett. 56, 930–933 (1986).
2. G. Binnig, H. Rohrer, Ch. Gerber & E. Weibel, "Surface studies by Scanning Tunneling Microscopy", Phys. Rev. Lett. 49 57–61 (1982).
HS-AFM (High Speed Atomic Force Microscopy)
タンパク質の大きさは通常10~50 nm(ナノメートル)程度で、通常の顕微鏡ではその構造や動態を直接観察することはできません。これまで光学顕微鏡やX線結晶構造解析などが用いられてきましたが、溶液中での動態観察には限界がありました。一方、AFMは「触覚」の原理を応用して、針と試料の間に働く微小な力を検出することで表面構造を画像化します。この特性により、AFMは溶液環境下で動作可能で、生きたタンパク質をナノスケールで観察できるという利点を持っています。しかし従来のAFMでは画像化速度が遅く、動的な分子挙動をリアルタイムで観察することは困難でした。当研究室では、AFMの高速化に取り組み、1枚の画像を100ミリ秒(ms)で取得できる「高速AFM」を開発しました。この技術により、タンパク質が溶液中で動く様子をリアルタイムで撮影でき、動的な分子機構の解明に大きく貢献しています。例えば、高速AFMを用いてアルツハイマー病の原因タンパク質であるアミロイドベータ(Aβ)の線維化過程を詳細に観察し、線維形成の初期過程や成長メカニズムを明らかにしました。また、線維形成を阻害する抗体との相互作用を観察することで、新たな治療法や創薬研究にも貢献しています。このように、高速AFMは基礎から応用まで幅広い分野に活用されています。
AFMの動作原理
アミロイド線維の伸長過程の観察
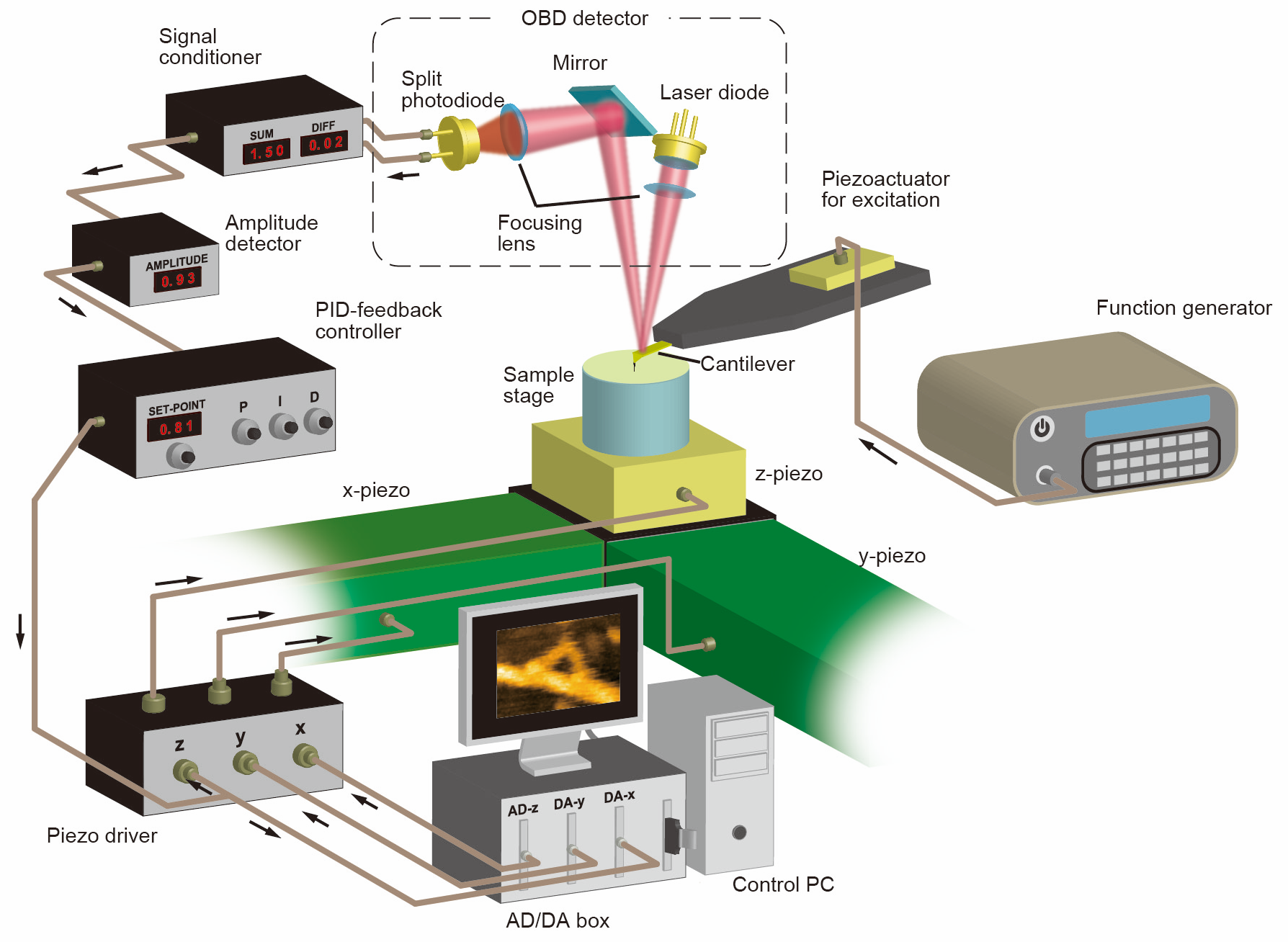
高速AFMの構成
アミロイド線維への抗体の結合
HS-AFM (High Speed Atomic Force Microscopy)
Although several operation modes have been proposed for AFM, a method called tapping mode is often used for soft samples such as biomolecules. The high-speed AFM also emplopys the tapping mode. In the tapping mode, the cantilever with the probe is oscillated at the mechanical resonance frequency by a piezoelectric actuator and the change in the oscillation amplitude of the cantilever wwhen the probe intermittently contacts the sample is detected by theoptical lever method. While the probe is raster-scanned against the sample, the motion of piezoelectric actuator in the Z direction is controlled by PID feedback control so that the oscillation amplitude of the cantilever is kept to be the reference value, and the distance between the probe and the sample is kept constant. The PID signal at each pixel position of the two-dimensional scanning can be recorded by a computer to image a surface profile. To increase the frame rate of AFM, all the devices composing AFM must be operated at high speed. Several research groups have attempted to improve the temporal resolution of AFM and to observe the dynamics of biological samples. In 1989, it was reported that the polymerization process of fibrinogen, one of the blood coagulation factors, was observed at an imaging rate of 8 s/frame. In the 1990s, many studies of AFM observation of biomolecules at imaging speeds of a few seconds were reported, but no group has succeeded in imaging fragile biomolecules on the time scale in which they function (typically, less than 1 second) without damaging their functions. Prof. Toshio Ando of Kanazawa University started to improve the speed of AFM around 1993 with the goal of developing an instrument that could observe the structural dynamics of proteins, and in 2001 succeeded in imaging the dynamics of proteins adsorbed on a mica substrate at 80 ms/frame [2].Prof. Uchihashi joined Prof. Ando’s group in 2004 and has been working on the improvement and application of high-speed AFM. As a result of technological improvements in high-speed AFM, such as minimization of invasiveness, higher resolution, and ease of operation, a system capable of high-speed imaging of dynamics such as protein’s structural dynamcics, molecular diffusion and assembly processes was established around 2008 [3]. Since then, many studies using high-speed AFM have been conducted around the world [4].
1. B. Drake B, CB Prater, AL Weisenhorn, SA Gould, TR Albrecht, CF Quate, DS Cannell, HG Hansma & PK Hansma, "Imaging crystals, polymers, and processes in water with the atomic force microscope" Science Science 24 , 1586–1589 (1989).
2. T. Ando, N. Kodera, E. Takai, D. Maruyama, K. Saito & A. Toda, "A high-speed atomic force microscope for studying biological macromolecules", Proc. Natl. Acad. Sci. USA. 98, 12468–12472 (2001).
3. T. Ando, T. Uchihashi & T. Fukuma, "High-speed atomic force microscopy for nano-visualization of dynamic biomolecular processes", Prog. Surf. Sci. 83, 337–437 (2008).
4. T. Ando, T. Uchihashi & S. Scheuring, "Filming biomolecular processes by high-speed atomic force microscopy", Chem. Rev. 114, 3120–3188 (2014).
大腸菌の分裂過程における形状と硬さの時間変化の観察
bottom of page

